Usage
This section describes the core usage of the colabfit-tools package.
Parsing data
Properties can be added into the Database is by creating or loading a
Configuration object, attaching the Property data to the info
and/or arrays dictionaries of the Configuration (see here for more details), then using the
insert_data() method.
In addition to the Configuration with attached Property information,
insert_data() will also require a
property map describing how to map the attached Property information onto an
existing property definition (see Property definitions).
A property map should have the following structure:
{
<property_name>: {
<property_field>: {
'field': <key_for_info_or_arrays>,
'units': <ase_readable_units>
}
}
}
See below for the definitions of each of the above keys/values:
<property_name>should be one of the following:The name of an OpenKIM Property Definition from the list of approved OpenKIM Property Definitions
The name of a locally-defined property (see Property definitions) that has been added using
insert_property_definition()
<property_field>should be the name of a field from an property definition<key_for_info_or_arrays>should be a key for indexing theinfoorarraysdictionaries on a Configuration (see Configuration info and arrays fields)'field'is used to specify the key for extracting the property fromConfiguration.infoorConfiguartion.arrays.'units'should be a string matching one of the units names in ase.units.
Note that insert_data() will attempt to load every Property specified in
property_map for each Configuration. This means that if there are
P properties in property_map and C Configurations, a
maximum of P*C Properties will be loaded in total. If a Configuration does not
have the necessary data for loading a given Property, that Property is skipped
for the given Configuration and a warning is raised.
Detecting duplicates
All entities in the Database (Configuration, Property, PropertySetting,
ConfigurationSet, and Dataset) are stored with a unique ID that is generated by
hashing the corresponding entity (see the documentation for their respective
hash functions for more detail). Because of this, if two entities are identical,
they will only be stored once in the Database. This can be useful when trying to
remove duplicate data in a Dataset, as it ensures that no duplicates are added
during insert_data().
For example, this enables a user to take advantage of the Mongo
count_documents() function to see if a Configuration is linked to more
than one Property
client.configurations.count_documents(
{'relationships.properties.2': {'$exists': True}}
)
Similarly, the 'relationships' fields of entities can be used to check
if two Datasets have any common Configurations.
Synchronizing a Dataset
When working with a Dataset, it is important to make sure that the dataset has been “synchronized” in order to ensure that all of the data (configuration labels, configuration sets, aggregated metadata, …) have been properly updated to reflect any recent changes to the Dataset.
There are three points in the Database where data aggregation is performed (and thus where care must be taken to ensure synchronization):
ConfigurationSets aggregating Configuration information (
aggregate_configuration_info())Datasets aggregating ConfigurationSet information (
aggregate_configuration_set_info())Datasets aggregating Property information (
aggregate_property_info())
Synchronization can be performed by using the resync=True argument when
calling get_configuration_set() and
get_dataset(). Aggregation is
automatically performed when inserting ConfigurationSets. Aggregation is not
automatically performed when inserting Datasets in order to avoid re-aggregating
ConfigurationSet information unnecessarily; therefore resync=True may
also be used for insert_dataset().
In order to re-synchronize the entities without using the get_*() methods,
call the aggregate_*() methods directly. A common scenario where this may
be necessary is when using
apply_labels() in order to make
sure that the changes are reflected in the ConfigurationSets and Datasets.
Applying configuration labels
Configuration labels should be applied using the
apply_labels() method.
An example configuration_label_regexes:
client.apply_labels(
dataset_id=ds_id,
collection_name='configurations',
query={'nsites': {'$lt': 100}},
labels={'small'},
verbose=True
)
See the Si PRX GAP tutorial for a more complete example.
Building configuration sets
There are two steps to building a ConfigurationSet:
First, extracting the IDs of the Configurations that should be included in the ConfigurationSet:
co_ids = client.get_data(
'configurations',
fields='_id',
query={'_id': {'$in': <all_co_ids_in_dataset>}, 'nsites': {'$lt': 100}},
ravel=True
).tolist()
And second, calling
insert_configuration_set():
cs_id = client.insert_configuration_set(
co_ids,
description='Configurations with fewer than 100 atoms'
)
Note that in the first step, '_id': {'$in': <all_co_ids_in_dataset>} is
used in order to limit the query to include only the Configurations within a
given Dataset, rather than all of the Configurations in the entire Database.
See the Si PRX GAP tutorial for a more complete example.
Attaching property settings
A PropertySettings object can be
attached to a Property by specifying the property_settings argument
in insert_data().
pso = PropertySettings(
method='VASP',
description='A basic VASP calculation',
files=None,
labels=['PBE', 'GGA'],
)
client.insert_data(
images,
property_map=...,
property_settings={
<property_name>: pso,
}
)
Data exploration
The get_data(),
plot_histograms(),
and get_statistics() functions can
be extremely useful for quickly visualizing your data and detecting outliers.
energies = client.get_data('properties', '<property_name>.energy', ravel=True)
forces = client.get_data('properties', '<property_name>.forces', concatenate=True)
# From the QM9 example
client.get_statistics(
['qm9-property.a', 'qm9-property.b', 'qm9-property.c'],
ids=dataset.property_ids,
verbose=True
)
client.plot_histograms(
['qm9-property.a', 'qm9-property.b', 'qm9-property.c',],
ids=dataset.property_ids
)
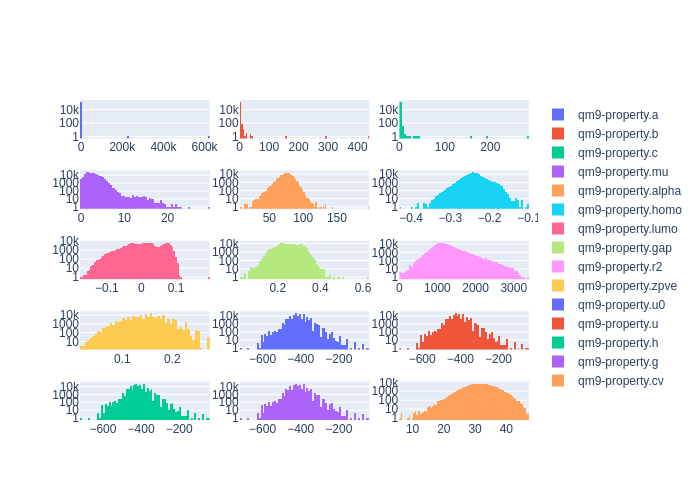
See the QM9 example and the Si PRX GAP example to further explore the benefits of these functions.
Since Configurations inherit from ase.Atoms objects, they work
seamlessly with ASE’s visualization tools.
# Run inside of a Jupyter Notebook
configurations = client.get_configurations(configuration_ids)
from ase.visualize import view
# Creates a Jupyter Widget; may require `pip install nglview` first
view(configurations, viewer='nglview')
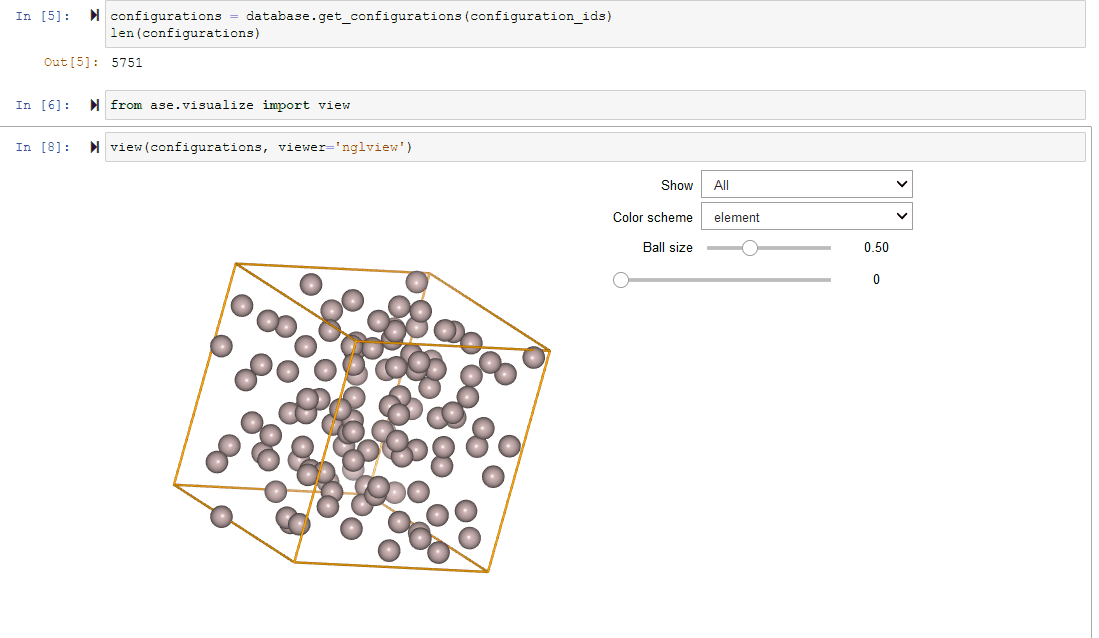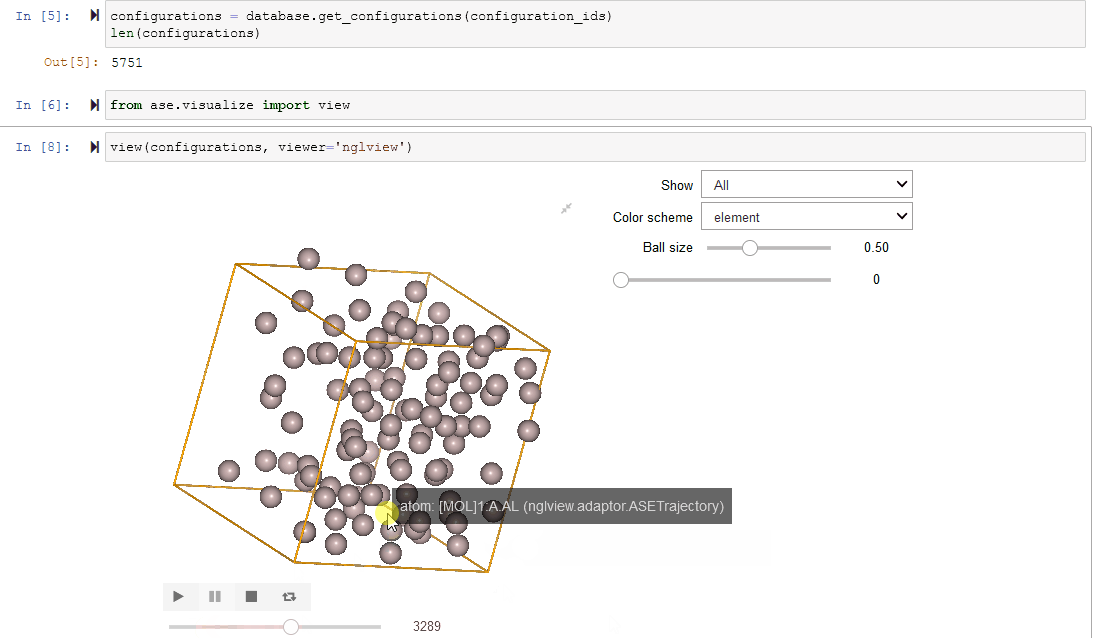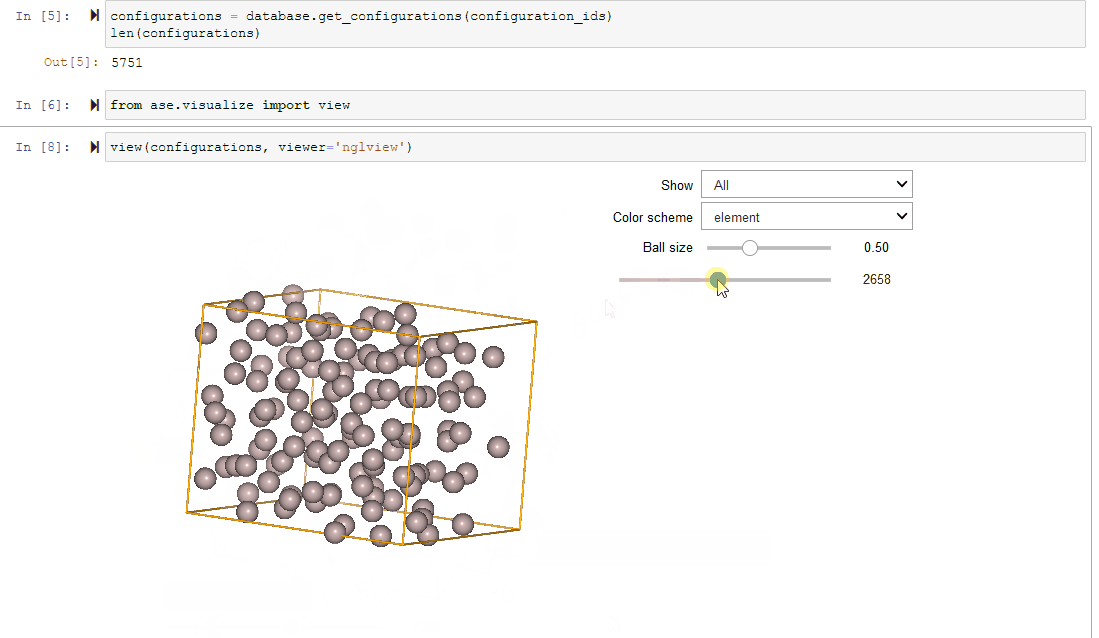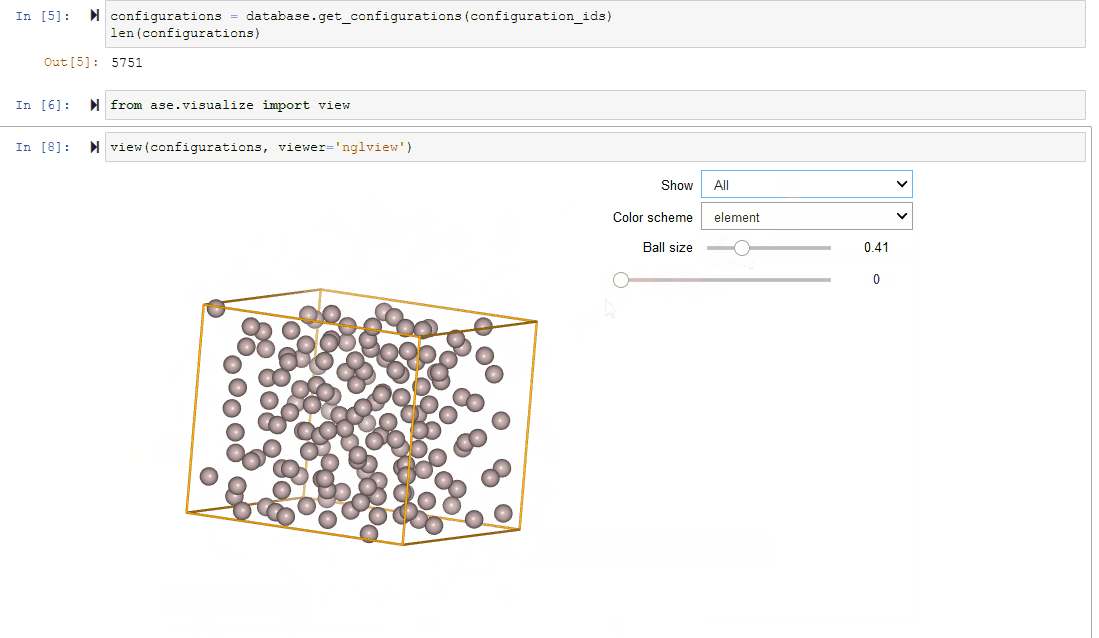
Filtering a Dataset
Datasets can be easily filtered to remove unwanted entries or extract subsets of
interest. Filtering can be done using the
filter_on_properties() or
filter_on_configurations()
methods.
# From the QM9 example
clean_config_sets, clean_property_ids = client.filter_on_properties(
ds_id=ds_id,
filter_fxn=lambda x: (x['qm9-property']['a']['source-value'] < 20)
and x['qm9-property']['b']['source-value'] < 10,
fields=['qm9-property.a.source-value', 'qm9-property.b.source-value'],
verbose=True
)
Note the use of the ds_id argument, which makes sure that the returned
ConfigurationSet IDs and Property IDs are only those that are contained within
the given Dataset.
Data transformations
It is often necessary to transform the data in a Dataset in order to improve
performance when fitting models to the data, or to convert the data into a
different format. This can be done using the transfom argument of the
insert_data() function. The transform argument
should be a callable function that modifies the Configuration in-place:
def per_atom(c):
c.info['energy'] /= len(c)
client.insert_data(
configurations,
property_map=property_map,
property_settings=property_settings,
transform=per_atom,
)
Supported file formats
Ideally, raw data should be stored in Extended XYZ format. This is the
default format used by colabfit-tools, and should be suitable for almost
all use cases. CFG files (used by Moment Tensor Potentials) are also supported,
but are not recommended.
Data that is in a custom format (e.g., JSON, HDF5, …) that cannot be easily
read by
ase.io.read will
require the use of a FolderConverter
instance, which needs to be supplied with a custom reader() function for
parsing the data.