Database structure
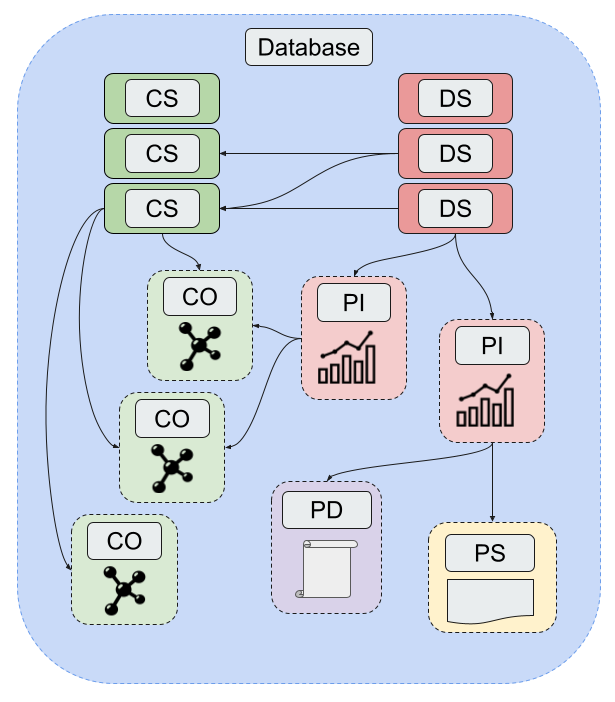
A diagram showing the relationship between the core components of a Database.
The Database structure was designed to be able to be as flexible as possible to incorporate different types of data (computational/experimental) while also making the datasets efficient to query, store, and manipulate. A Database is stored as a Mongo database with five core data structures:
Configuration(CO,collection_name='configurations'):The information necessary to uniquely define the input to a material property calculation or the atomic geometry present in an experimental measurement. At a minimum, a configuration must include the atomic species and nuclear positions. In the case of periodic or semi-periodic systems, the simulation cell vectors must also be included. Information related to atomic charges, magnetic moments, and electric dipoles/quadrupoles may optionally be specified, and may serve either as constraints on a first-principles model or as inputs to an effective model. Additional metadata related to the configuration can also be provided (e.g., the parent structure from which it was generated by perturbing its positions).
Property(PI,collection_name='properties'):The outputs from a material property calculation, e.g. DFT-computed energy/forces/stress, or experimental measurement. A property instance points to one or more individual
Configurationinstances; if theConfigurationobjects that a property points to contain optional inputs such as charges or magnetic moments, then the property must contain an associated output value for each of them (in the event that these inputs serve as constraints, the output values will be equal to the input values). Generally, it is best practice for a property to point to aPropertySettingsobject.
Property Definition(PD,collection_name='property_definitions'):A Python dictionary that specifies details about the contents of a Property, including the name of the property type, a description of the property, and information about each of the computed fields in the property (data type, data shape, if the field has units, if the field is required, and a human-readable description of the field).
PropertySettings(PS,collection_name='property_settings'):Additional metadata useful for setting up the calculation or experiment (e.g., the name of the software package(s) used, their versions, input files, experimental method or devices, etc.).
ConfigurationSet(CS,collection_name='configuration_sets'):An object defining a group of one or more
Configurationinstances and providing useful metadata for organizing datasets (e.g., “Snapshots from a molecular dynamics run at 1000K”).
Dataset(DS,collection_name='datasets'):An object used to effectively aggregate information from all of the data structures defined above to create a body of information that provides a complete, discoverable training set. A
Datasetpoints to one or moreConfigurationSetobjects, one or morePropertyobjects, and one or more otherDatasetobjects.